Discovery and development of NS5A inhibitors
 From Wikipedia the free encyclopedia
From Wikipedia the free encyclopedia

Nonstructural protein 5A (NS5A) inhibitors are direct acting antiviral agents (DAAs) that target viral proteins, and their development was a culmination of increased understanding of the viral life cycle combined with advances in drug discovery technology.[1][2] However, their mechanism of action is complex and not fully understood.[2] NS5A inhibitors were the focus of much attention when they emerged as a part of the first curative treatment for hepatitis C virus (HCV) infections in 2014.[3] Favorable characteristics have been introduced through varied structural changes, and structural similarities between NS5A inhibitors that are clinically approved are readily apparent.[4][5] Despite the recent introduction of numerous new antiviral drugs, resistance is still a concern and these inhibitors are therefore always used in combination with other drugs.[6][7]
Hepatitis C virus
[edit]HCV is a positive-sense single-stranded RNA virus that has been demonstrated to replicate in the hepatocytes of both humans and chimpanzees. A single HCV polyprotein is translated, and then cleaved by cellular and viral proteases into three structural proteins (core, E1, and E2) and seven nonstructural proteins (p7, NS2, NS3, NS4A, NS4B, NS5A, and NS5B).[8][9]
HCV is among the leading causes of liver disease around the world. It is transmitted by blood and is most commonly contracted through the use of infected needles.[10] Patients with chronic HCV infection are at significant risk of cirrhosis and hepatocellular carcinoma, which are the leading causes of death for those infected.[11][12]
The virus has been around for over a millennium and has been classified into six known genotypes, each of which contains numerous subtypes. The seventh remains uncharacterized. The genotype contracted dictates which specific treatments are viable.[13]
NS5A receptor
[edit]Basic structure and chemical properties
[edit]NS5A is a large hydrophilic phosphoprotein that is essential for the HCV life cycle and is found in association with virus-induced membrane vesicles, termed the membranous web.[14][15] NS5A is a proline-rich protein composed of approximately 447 amino acids, which is divided into three domains.[16][17] These domains are linked by two low-complexity sequences that are either serine- or proline-rich.[18] Domain I is a zinc binding domain and X-ray crystallography studies indicated alternative dimer conformations of domain I of NS5A.[19][20][21] Domain II and III are unstructured, shown by NMR studies.[16][22] Domain I is preceded by an N-terminal amphipathic helix which allows the protein to associate with endoplasmic reticulum-derived membranes.[22][23][24] Although X-ray crystallographic studies revealed dimer conformations of NS5A domain1, recent in solution structural characterization studies showed that NS5A proteins form higher-order structures by dimeric subunits of NS5A domain 1.[25] Moreover, the overall structural model of NS5A highlights the variability of intrinsic conformations of the D2 and D3 domains between HCV genotypes.[26] Therefore, it is still under debate which conformation\s of NS5A is functional and also targeted by NS5A inhibitors.[citation needed]
NS5A mainly exists in two distinct phosphorylated forms, a hypophosphorylated and a hyperphosphorylated form, but the exact function of the phosphorylation has not been determined.[17][18][27]
Function
[edit]The NS5A protein plays an important role in viral RNA replication, viral assembly, and complex interactions with cellular functions.[2][17] The protein has been implicated in the modulation of host defenses, apoptosis, the cell cycle, and stress-responsive pathways.[27] However, its function and complete structure have yet to be elucidated.[16]
NS5A seems to be key in triggering the formation of the membranous web in the absence of other similar nonstructural proteins.[15] Many proteins within the host cell can be affected by NS5A, e.g. phosphatidylinositol 4-kinase IIIα (PI4KIIIα), a kinase required for the replication of HCV. This kinase takes part in the biosynthesis of phosphatidylinositol 4-phosphate (PI4P) by interacting with NS5A, which stimulates its activity and appears to improve the integrity of the membranous web.[15][28][29]
Recently, the central role of NS5A in viral proliferation has made it the target for drug development. As a result, new antiviral agents have been introduced for the treatment of HCV.[2]
Mechanism of action
[edit]NS5A inhibitors have been developed to target the NS5A protein. These inhibitors have achieved a significant reduction in HCV RNA blood levels and can therefore be considered as potent antivirals. Their mechanism of action is thought to be diverse but the exact mechanism is not fully understood.[2][30] Most studies assume that NS5A inhibitors act on two essential stages of the HCV life cycle; the replication of the genomic RNA, and virion assembly. Other studies propose an alteration of host cell factors as a possible third mechanism.[2][15][31]

The structure of NS5A inhibitors is characterized by dimeric symmetry. This suggests that NS5A inhibitors act on dimers of NS5A.[32] A number of modeling studies have shown that daclatasvir, which is an NS5A inhibitor, only binds to the "back-to-back" NS5A dimer and that the binding has to be symmetrical. Other modeling studies have shown that binding to other conformations of NS5A might be possible, as well as asymmetrical binding.[30] Research has shown that daclatasvir's target is most likely domain I of NS5A.[31] Even though the mechanism is not completely understood, it has been demonstrated that the inhibitors downregulate NS5A hyperphosphorylation, leading to the suppression of HCV replication and its processing of polyproteins, as well as resulting in an unusual protein location.[31][33] Hitherto, this inhibition was thought to require only NS5A domain I, but not domains II and III.[33] However, recent studies have shown that both domains I and II are relevant to this disruption of RNA replication.[34]
NS5A inhibitors appear to furthermore disrupt the formation of new replicase complexes resulting in a gradual slowing of viral RNA synthesis. Effect on previously formed complexes has yet to be demonstrated.[34][35]
Available evidence suggests that NS5A inhibitors modify the location of NS5A inside the cell. This may cause abnormal assembly leading to malformed viruses.[2] Some studies have revealed that inhibition of the viral assembly has a more important role in RNA reduction than viral replication reduction.[35][36]
Studies have shown that NS5A inhibitors block the formation of the membranous web, which protects the viral genome and features the main sites for viral replication and assembly.[15][31][37] This mechanism is thought to be independent of RNA replication, but seems to be affected by NS5A inhibitors blocking the formation of the PI4KIIIα-NS5A complex, essential to the synthesis of the PI4P, resulting in decreased integrity of the membranous web and therefore reduced HCV RNA replication.[15][28][38]
History
[edit]HCV research has taken great strides in recent years with the discovery and clinical development of multiple new HCV drugs. Among those drugs are the DAAs which include NS5A inhibitors.[39] NS5A inhibitors have been found particularly effective in the treatment of HCV where they have been used in combination with protease inhibitors such as glecaprevir or NS5B inhibitors (e.g., sofosbuvir), pegylated interferons (e.g. peginterferon alfa-2a), and ribonucleic analogs (e.g. ribavirin).[40][41][42] The ever present risk of viral strains developing resistance has been a main factor in why they are used in combination with one or more complementary drug.[43]
Adverse effects, and extensive and complicated drug regimens with accompanying low compliance rates, have been a hindrance in the development of antiviral treatments. The combination of NS5A and NS5B inhibitors has produced positive results in this regard.[44]
Drug discovery and development
[edit]Discovery
[edit]
The discovery of NS5A inhibitors took place within the context of a pursuit for a treatment for HCV. NS5A is among the seven nonstructural proteins that form a complex with viral RNA within infected cells to initiate HCV replication.[45] HCV research has produced several DAAs including NS3A, NS4A and NS5B inhibitors, as well as NS5A inhibitors.[46]
Development
[edit]The development of antiviral drugs capable of interfering with the proteins responsible for viral replication has been intimately linked with advancements in techniques for establishing the efficient cell culture systems needed to screen for them.[46]
In 1999 a breakthrough came when a full-length consensus genome cloned from HCV RNA was found to replicate at high levels when transfected into a human hepatoma cell line.[47] This method has since been improved upon with the use of cell culture-adaptive mutations that enhance RNA replication.[48]
Screening has now produced a number of NS5A inhibitors, which have been incorporated into treatments for HCV. The first in this new class of drugs was daclatasvir (Daklinza), gaining first global approval from the Japanese Ministry of Health, Labour and Welfare (MHLW) in July 2014 in combination with asunaprevir.[49] Daclatasvir received FDA approval in July 2015.[50] Other drugs have since been approved, among them notably the first FDA-approved NS5A inhibitor ledipasvir, approved October 2014 in combination with sofosbuvir to comprise the HCV drug Harvoni.[51][52]
Although NS5A inhibitors have proven effective antivirals, they must be used alongside complementary antiviral drugs due to how quickly they lead to the development of resistant mutations when given as a single agent.[53] This has shaped the focus of NS5A inhibitor development, from which asymmetrical variants that metabolize into analogues with complementary resistance profiles have emerged, amongst other discoveries.[54]
Structure-activity relationship
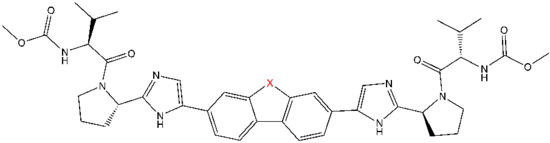
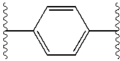
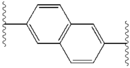
[edit]The structural similarities between the inhibitors are readily apparent.[4] The appendages of the central core are typically symmetrical and have an imidazole-proline structure.[5] The natural L-configuration of the proline derivatives was found to be critical for inhibition since the unnatural D-configuration had drastically weaker activity. The potency of the inhibitors was correspondingly sensitive to changes in the amine capping element. These observations suggest that the amine region of the molecules plays an important role in the inhibitory activity.[55]
Favorable characteristics in an NS5A inhibitor include high potency and long plasma half-life in order to achieve a once-daily-dosage. Slightly asymmetrical appendages, as seen in ledipasvir, were found to have distinctive benefits for the optimization of inhibitor potency and pharmacokinetics.[51] The structure of the central core changes the spacing and the projection of the appendages as well as the position of the lipophilicity in the central core, which affects inhibitory activity notably. Structures with fused central rings consistently show greater inhibitory activity, whereas less lipophilic central cores provide weaker activity.[4] Symmetrical bis-imidazol structures, such as daclatasvir, experience a loss in potency when fluorene is substituted for the biaryl group. This substitution also gives rise to some serious stability problems.[51][56] However, a smaller lipophilic connector such as difluoromethylene generates the most potent inhibitor in an asymmetrical structure. Additionally, it provides improved bioavailability and more favorable plasma half-life. There is also a remarkable increase in potency when phenyl is replaced with naphthyl as a central core. This increase is significantly higher in an asymmetrical structure than it is in a symmetrical structure.[51][55] In asymmetrical structures, a difference in potency between the phenyl-alkyne inhibitors demonstrates the importance of the position of the lipophilicity. A more centrally located alkyne, which is a less lipophilic connector than phenyl, improves potency.[4][5][51]
| Structure | Activity | |
|---|---|---|
| X | IC50 (nM) | Inhibitory activity |
| >44 | None | |
| >44 | None | |
| 11 | Very weak | |
| 1.7 | Weak | |
| 0.50 | Moderate | |
| 3.7 | Weak | |
| 0.11 | Moderate | |
| 0.20 | Moderate | |
| Structure | Activity | |
|---|---|---|
| X | IC50 (nM) | Inhibitory activity |
| >44 | None | |
| 0.071 | Moderate | |
| 2.5 | Weak | |
| 0.38 | Moderate | |
| 0.20 | Moderate | |
| 0.17 | Moderate | |
| 0.040 | Strong | |
| Structure | Activity | |
|---|---|---|
| X | IC50 (nM) | Inhibitory activity |
| CH2 | 0.094 | Strong |
| CO | 0.30 | Moderate |
| C(CH3)2 | 1.2 | Weak |
Resistance
[edit]The potential HCV resistance against DAA drugs is a concern.[6] Among the HCV quasispecies there are pre-existing variants with the potential to confer resistance to NS5A inhibitors without having any previous exposure to those drugs. Generally, the replication of these variants happens only in minute quantities, making them undetectable by current techniques. On the other hand, it is possible to selectively grow immune variants in the presence of NS5A inhibitors.[2] HCV resistance is characterized by a certain escape pattern. This pattern is often associated with amino acid substitutions that confer upon the virus a robust drug resistance without impairing the viral fitness.[2][57] It has been established that NS5A inhibitors possess a relatively low threshold for resistance, and variants that are associated with NS5A resistance have been shown to endure for up to six months in patients following treatment cessation.[58] Therefore, combination therapies produce higher efficacy and shorter treatment periods.[7]
Future research and new generations of NS5A inhibitors
[edit]DAA developers face foreseeable challenges in the years to come. Therapeutic gaps for individuals with complicating conditioned such as chronic kidney disease and cirrhosis will need to be bridged. Shorter therapies with milder side effects would yield greater adherence, and the ever present spectre of drug resistance is looming. The highly adaptive HCV has evolved into a number of different genomes that all need to be adequately treated, preferably with pan-genotypic regimens.[59]
Some of these challenges already have possible solutions in sight. The protease inhibitor ABT-493 and the next-generation NS5A inhibitor ABT-530 are considered active against all HCV genotypes, including the hard to treat genotype 3.[42][59] In vitro, ABT-530 showed potency against the resistance associated variants which are immune to the first generations of NS5A inhibitors, including ledipasvir, daclatasvir and ombitasvir.[42] Because this drug combination has the additional quality of being hepatically cleared, it holds the promise that patients with chronic kidney disease and HCV could receive a safe, non-sofosbuvir-based treatment in the near future.[59]
At least three drug combinations for the treatment of HCV are in the pipeline to be approved in 2016-2017: Sofosbuvir in combination with velpatasvir, ABT-493 in combination with ABT-530, and grazoprevir in combination with elbasvir, of which velpatasvir, ABT-530 and elbasvir are NS5A inhibitors.[7]
See also
[edit]References
[edit]- ^ Gogela, Neliswa A.; Lin, Ming V.; Wisocky, Jessica L.; Chung, Raymond T. (12 March 2015). "Enhancing Our Understanding of Current Therapies for Hepatitis C Virus (HCV)". Current HIV/AIDS Reports. 12 (1): 68–78. doi:10.1007/s11904-014-0243-7. PMC 4373591. PMID 25761432.
- ^ a b c d e f g h i Pawlotsky, Jean-Michel (August 2013). "NS5A inhibitors in the treatment of hepatitis C". Journal of Hepatology. 59 (2): 375–382. doi:10.1016/j.jhep.2013.03.030. PMID 23567084.
- ^ Do, Albert; Mittal, Yash; Liapakis, AnnMarie; Cohen, Elizabeth; Chau, Hong; Bertuccio, Claudia; Sapir, Dana; Wright, Jessica; Eggers, Carol; Drozd, Kristine; Ciarleglio, Maria; Deng, Yanhong; Lim, Joseph K.; Jhaveri, Ravi (27 August 2015). "Drug Authorization for Sofosbuvir/Ledipasvir (Harvoni) for Chronic HCV Infection in a Real-World Cohort: A New Barrier in the HCV Care Cascade". PLOS ONE. 10 (8): e0135645. Bibcode:2015PLoSO..1035645D. doi:10.1371/journal.pone.0135645. PMC 4552165. PMID 26312999.
- ^ a b c d e f g h Tong, Ling; Yu, Wensheng; Coburn, Craig A.; Meinke, Peter T.; Nair, Anilkumar G.; Dwyer, Michael P.; Chen, Lei; Selyutin, Oleg; Rosenblum, Stuart B.; Jiang, Yueheng; Fells, James; Hu, Bin; Zhong, Bin; Soll, Richard M.; Liu, Rong; Agrawal, Sony; Xia, Ellen; Zhai, Ying; Kong, Rong; Ingravallo, Paul; Nomeir, Amin; Asante-Appiah, Ernest; Kozlowski, Joseph A. (July 2016). "Alternative core development around the tetracyclic indole class of HCV NS5A inhibitors". Bioorganic & Medicinal Chemistry Letters. 26 (20): 5132–5137. doi:10.1016/j.bmcl.2016.07.057. PMID 27634194.
- ^ a b c Lemm, J. A.; Leet, J. E.; O'Boyle, D. R.; Romine, J. L.; Huang, X. S.; Schroeder, D. R.; Alberts, J.; Cantone, J. L.; Sun, J.-H.; Nower, P. T.; Martin, S. W.; Serrano-Wu, M. H.; Meanwell, N. A.; Snyder, L. B.; Gao, M. (16 May 2011). "Discovery of Potent Hepatitis C Virus NS5A Inhibitors with Dimeric Structures". Antimicrobial Agents and Chemotherapy. 55 (8): 3795–3802. doi:10.1128/AAC.00146-11. PMC 3147613. PMID 21576451.
- ^ a b Fridell, R. A.; Qiu, D.; Wang, C.; Valera, L.; Gao, M. (28 June 2010). "Resistance Analysis of the Hepatitis C Virus NS5A Inhibitor BMS-790052 in an In Vitro Replicon System". Antimicrobial Agents and Chemotherapy. 54 (9): 3641–3650. doi:10.1128/AAC.00556-10. PMC 2935007. PMID 20585111.
- ^ a b c Asselah, Tarik; Boyer, Nathalie; Saadoun, David; Martinot-Peignoux, Michele; Marcellin, Patrick (January 2016). "Direct-acting antivirals for the treatment of hepatitis C virus infection: optimizing current IFN-free treatment and future perspectives". Liver International. 36: 47–57. doi:10.1111/liv.13027. PMID 26725897.
- ^ Halliday, John; Klenerman, Paul; Barnes, Eleanor (9 January 2014). "Vaccination for hepatitis C virus: closing in on an evasive target". Expert Review of Vaccines. 10 (5): 659–672. doi:10.1586/erv.11.55. PMC 3112461. PMID 21604986.
- ^ Grakoui, A; Wychowski, C; Lin, C; Feinstone, S M; Rice, C M (1 March 1993). "Expression and identification of hepatitis C virus polyprotein cleavage products". Journal of Virology. 67 (3): 1385–1395. doi:10.1128/JVI.67.3.1385-1395.1993. ISSN 0022-538X. PMC 237508. PMID 7679746.
- ^ Chan, Juliana (May 2014). "Hepatitis C". Disease-a-Month. 60 (5): 201–212. doi:10.1016/j.disamonth.2014.04.002. PMID 24863270.
- ^ Seeff, Leonard B. (November 2002). "Natural history of chronic hepatitis C". Hepatology. 36 (5): s35 – s46. doi:10.1053/jhep.2002.36806. PMID 12407575.
- ^ Liang, T. Jake; Rehermann, Barbara; Seeff, Leonard B.; Hoofnagle, Jay H. (15 February 2000). "Pathogenesis, Natural History, Treatment, and Prevention of Hepatitis C". Annals of Internal Medicine. 132 (4): 296–305. doi:10.7326/0003-4819-132-4-200002150-00008. PMID 10681285. S2CID 23226526.
- ^ Pybus, O. G.; Barnes, E.; Taggart, R.; Lemey, P.; Markov, P. V.; Rasachak, B.; Syhavong, B.; Phetsouvanah, R.; Sheridan, I.; Humphreys, I. S.; Lu, L.; Newton, P. N.; Klenerman, P. (29 October 2008). "Genetic History of Hepatitis C Virus in East Asia". Journal of Virology. 83 (2): 1071–1082. doi:10.1128/JVI.01501-08. PMC 2612398. PMID 18971279.
- ^ Tellinghuisen, T. L.; Foss, K. L.; Treadaway, J. C.; Rice, C. M. (21 November 2007). "Identification of Residues Required for RNA Replication in Domains II and III of the Hepatitis C Virus NS5A Protein". Journal of Virology. 82 (3): 1073–1083. doi:10.1128/JVI.00328-07. PMC 2224455. PMID 18032500.
- ^ a b c d e f Reghellin, V.; Donnici, L.; Fenu, S.; Berno, V.; Calabrese, V.; Pagani, M.; Abrignani, S.; Peri, F.; De Francesco, R.; Neddermann, P. (15 September 2014). "NS5A Inhibitors Impair NS5A-Phosphatidylinositol 4-Kinase III Complex Formation and Cause a Decrease of Phosphatidylinositol 4-Phosphate and Cholesterol Levels in Hepatitis C Virus-Associated Membranes". Antimicrobial Agents and Chemotherapy. 58 (12): 7128–7140. doi:10.1128/AAC.03293-14. PMC 4249536. PMID 25224012.
- ^ a b c Yamasaki, Lilian HT; Arcuri, Helen A; Jardim, Ana Carolina G; Bittar, Cintia; de Carvalho-Mello, Isabel Maria VG; Rahal, Paula (2012). "New insights regarding HCV-NS5A structure/function and indication of genotypic differences". Virology Journal. 9 (1): 14. doi:10.1186/1743-422X-9-14. PMC 3271958. PMID 22239820.
- ^ a b c Masaki, T.; Matsunaga, S.; Takahashi, H.; Nakashima, K.; Kimura, Y.; Ito, M.; Matsuda, M.; Murayama, A.; Kato, T.; Hirano, H.; Endo, Y.; Lemon, S. M.; Wakita, T.; Sawasaki, T.; Suzuki, T. (23 April 2014). "Involvement of Hepatitis C Virus NS5A Hyperphosphorylation Mediated by Casein Kinase I- in Infectious Virus Production". Journal of Virology. 88 (13): 7541–7555. doi:10.1128/JVI.03170-13. PMC 4054430. PMID 24760886.
- ^ a b Ross-Thriepland, D.; Harris, M. (20 November 2013). "Insights into the Complexity and Functionality of Hepatitis C Virus NS5A Phosphorylation". Journal of Virology. 88 (3): 1421–1432. doi:10.1128/JVI.03017-13. PMC 3911623. PMID 24257600.
- ^ Rice, Charles M.; Marcotrigiano, Joseph; Tellinghuisen, Timothy L. (May 2005). "Structure of the zinc-binding domain of an essential component of the hepatitis C virus replicase". Nature. 435 (7040): 374–379. Bibcode:2005Natur.435..374T. doi:10.1038/nature03580. ISSN 1476-4687. PMC 1440517. PMID 15902263.
- ^ Cronin, Ciarán N.; Wells, Peter A.; Hickey, Michael J.; Brodsky, Oleg; Love, Robert A. (2009-05-01). "Crystal Structure of a Novel Dimeric Form of NS5A Domain I Protein from Hepatitis C Virus". Journal of Virology. 83 (9): 4395–4403. doi:10.1128/JVI.02352-08. ISSN 1098-5514. PMC 2668466. PMID 19244328.
- ^ Lambert, Sebastian M.; Langley, David R.; Garnett, James A.; Angell, Richard; Hedgethorne, Katy; Meanwell, Nicholas A.; Matthews, Steve J. (2014-06-01). "The crystal structure of NS5A domain 1 from genotype 1a reveals new clues to the mechanism of action for dimeric HCV inhibitors". Protein Science. 23 (6): 723–734. doi:10.1002/pro.2456. ISSN 1469-896X. PMC 4093949. PMID 24639329.
- ^ a b Foster, T. L.; Belyaeva, T.; Stonehouse, N. J.; Pearson, A. R.; Harris, M. (30 June 2010). "All Three Domains of the Hepatitis C Virus Nonstructural NS5A Protein Contribute to RNA Binding". Journal of Virology. 84 (18): 9267–9277. doi:10.1128/JVI.00616-10. PMC 2937630. PMID 20592076.
- ^ Fridell, R. A.; Qiu, D.; Valera, L.; Wang, C.; Rose, R. E.; Gao, M. (18 May 2011). "Distinct Functions of NS5A in Hepatitis C Virus RNA Replication Uncovered by Studies with the NS5A Inhibitor BMS-790052". Journal of Virology. 85 (14): 7312–7320. doi:10.1128/JVI.00253-11. PMC 3126594. PMID 21593143.
- ^ a b Ascher, David B.; Wielens, Jerome; Nero, Tracy L.; Doughty, Larissa; Morton, Craig J.; Parker, Michael W. (23 April 2014). "Potent hepatitis C inhibitors bind directly to NS5A and reduce its affinity for RNA". Scientific Reports. 4: 4765. Bibcode:2014NatSR...4.4765A. doi:10.1038/srep04765. PMC 3996483. PMID 24755925.
- ^ Beldar, Serap; Manimekalai, Malathy Sony Subramanian; Cho, Nam-Joon; Baek, Kwanghee; Grüber, Gerhard; Yoon, Ho Sup (2018). "Self-association and conformational variation of NS5A domain 1 of hepatitis C virus". Journal of General Virology. 99 (2): 194–208. doi:10.1099/jgv.0.001000. PMID 29300159.
- ^ Badillo, Aurelie; Receveur-Brechot, Véronique; Sarrazin, Stéphane; Cantrelle, François-Xavier; Delolme, Frédéric; Fogeron, Marie-Laure; Molle, Jennifer; Montserret, Roland; Bockmann, Anja (2017-06-07). "Overall Structural Model of NS5A Protein from Hepatitis C Virus and Modulation by Mutations Confering [sic] Resistance of Virus Replication to Cyclosporin A". Biochemistry. 56 (24): 3029–3048. doi:10.1021/acs.biochem.7b00212. PMID 28535337.
- ^ a b Xiong, Wei; Yang, Jie; Wang, Mingzhen; Wang, Hailong; Rao, Zhipeng; Zhong, Cheng; Xin, Xiu; Mo, Lin; Yu, Shujuan; Shen, Chao; Zheng, Congyi; Diamond, M. S. (15 July 2015). "Vinexin β Interacts with Hepatitis C Virus NS5A, Modulating Its Hyperphosphorylation To Regulate Viral Propagation". Journal of Virology. 89 (14): 7385–7400. doi:10.1128/JVI.00567-15. PMC 4473562. PMID 25972535.
- ^ a b Reiss, Simon; Rebhan, Ilka; Backes, Perdita; Romero-Brey, Ines; Erfle, Holger; Matula, Petr; Kaderali, Lars; Poenisch, Marion; Blankenburg, Hagen; Hiet, Marie-Sophie; Longerich, Thomas; Diehl, Sarah; Ramirez, Fidel; Balla, Tamas; Rohr, Karl; Kaul, Artur; Bühler, Sandra; Pepperkok, Rainer; Lengauer, Thomas; Albrecht, Mario; Eils, Roland; Schirmacher, Peter; Lohmann, Volker; Bartenschlager, Ralf (January 2011). "Recruitment and Activation of a Lipid Kinase by Hepatitis C Virus NS5A Is Essential for Integrity of the Membranous Replication Compartment". Cell Host & Microbe. 9 (1): 32–45. doi:10.1016/j.chom.2010.12.002. PMC 3433060. PMID 21238945.
- ^ Lim, Y.-S.; Hwang, S. B. (5 February 2011). "Hepatitis C Virus NS5A Protein Interacts with Phosphatidylinositol 4-Kinase Type III and Regulates Viral Propagation". Journal of Biological Chemistry. 286 (13): 11290–11298. doi:10.1074/jbc.M110.194472. PMC 3064185. PMID 21297162.
- ^ a b Ahmed, Marawan; Pal, Abhishek; Houghton, Michael; Barakat, Khaled (3 August 2016). "A Comprehensive Computational Analysis for the Binding Modes of Hepatitis C Virus NS5A Inhibitors: The Question of Symmetry". ACS Infectious Diseases. 2 (11): 872–881. doi:10.1021/acsinfecdis.6b00113. PMID 27933783.
- ^ a b c d Issur, Moheshwarnath; Götte, Matthias (6 November 2014). "Resistance Patterns Associated with HCV NS5A Inhibitors Provide Limited Insight into Drug Binding". Viruses. 6 (11): 4227–4241. doi:10.3390/v6114227. PMC 4246218. PMID 25384189.
- ^ Lambert, Sebastian M.; Langley, David R.; Garnett, James A.; Angell, Richard; Hedgethorne, Katy; Meanwell, Nicholas A.; Matthews, Steve J. (June 2014). "The crystal structure of NS5A domain 1 from genotype 1a reveals new clues to the mechanism of action for dimeric HCV inhibitors". Protein Science. 23 (6): 723–734. doi:10.1002/pro.2456. PMC 4093949. PMID 24639329.
- ^ a b Qiu, D.; Lemm, J. A.; O'Boyle, D. R.; Sun, J.-H.; Nower, P. T.; Nguyen, V.; Hamann, L. G.; Snyder, L. B.; Deon, D. H.; Ruediger, E.; Meanwell, N. A.; Belema, M.; Gao, M.; Fridell, R. A. (27 July 2011). "The effects of NS5A inhibitors on NS5A phosphorylation, polyprotein processing and localization". Journal of General Virology. 92 (11): 2502–2511. doi:10.1099/vir.0.034801-0. PMID 21795470.
- ^ a b Belema, Makonen; Lopez, Omar D.; Bender, John A.; Romine, Jeffrey L.; St. Laurent, Denis R.; Langley, David R.; Lemm, Julie A.; O’Boyle, Donald R.; Sun, Jin-Hua; Wang, Chunfu; Fridell, Robert A.; Meanwell, Nicholas A. (13 March 2014). "Discovery and Development of Hepatitis C Virus NS5A Replication Complex Inhibitors". Journal of Medicinal Chemistry. 57 (5): 1643–1672. doi:10.1021/jm401793m. PMID 24621191.
- ^ a b Guedj, J.; Dahari, H.; Rong, L.; Sansone, N. D.; Nettles, R. E.; Cotler, S. J.; Layden, T. J.; Uprichard, S. L.; Perelson, A. S. (19 February 2013). "Modeling shows that the NS5A inhibitor daclatasvir has two modes of action and yields a shorter estimate of the hepatitis C virus half-life". Proceedings of the National Academy of Sciences. 110 (10): 3991–3996. Bibcode:2013PNAS..110.3991G. doi:10.1073/pnas.1203110110. PMC 3593898. PMID 23431163.
- ^ McGivern, David R.; Masaki, Takahiro; Williford, Sara; Ingravallo, Paul; Feng, Zongdi; Lahser, Frederick; Asante-Appiah, Ernest; Neddermann, Petra; De Francesco, Raffaele; Howe, Anita Y.; Lemon, Stanley M. (August 2014). "Kinetic Analyses Reveal Potent and Early Blockade of Hepatitis C Virus Assembly by NS5A Inhibitors". Gastroenterology. 147 (2): 453–462.e7. doi:10.1053/j.gastro.2014.04.021. PMC 4107048. PMID 24768676.
- ^ Neufeldt, Christopher J.; Joyce, Michael A.; Van Buuren, Nicholas; Levin, Aviad; Kirkegaard, Karla; Gale Jr., Michael; Tyrrell, D. Lorne J.; Wozniak, Richard W.; Kuhn, Richard J. (10 February 2016). "The Hepatitis C Virus-Induced Membranous Web and Associated Nuclear Transport Machinery Limit Access of Pattern Recognition Receptors to Viral Replication Sites". PLOS Pathogens. 12 (2): e1005428. doi:10.1371/journal.ppat.1005428. PMC 4749181. PMID 26863439.
- ^ Berger, Carola; Romero-Brey, Inés; Radujkovic, Danijela; Terreux, Raphael; Zayas, Margarita; Paul, David; Harak, Christian; Hoppe, Simone; Gao, Min; Penin, Francois; Lohmann, Volker; Bartenschlager, Ralf (November 2014). "Daclatasvir-Like Inhibitors of NS5A Block Early Biogenesis of Hepatitis C Virus–Induced Membranous Replication Factories, Independent of RNA Replication". Gastroenterology. 147 (5): 1094–1105.e25. doi:10.1053/j.gastro.2014.07.019. PMID 25046163.
- ^ a b Zhang, Xingquan (January 2016). "Direct anti-HCV agents". Acta Pharmaceutica Sinica B. 6 (1): 26–31. doi:10.1016/j.apsb.2015.09.008. PMC 4724659. PMID 26904396.
- ^ Fung, A.; Jin, Z.; Dyatkina, N.; Wang, G.; Beigelman, L.; Deval, J. (14 April 2014). "Efficiency of Incorporation and Chain Termination Determines the Inhibition Potency of 2'-Modified Nucleotide Analogs against Hepatitis C Virus Polymerase". Antimicrobial Agents and Chemotherapy. 58 (7): 3636–3645. doi:10.1128/AAC.02666-14. PMC 4068585. PMID 24733478.
- ^ Bruder Costa, Juliana; Dufeu-Duchesne, Tania; Leroy, Vincent; Bertucci, Inga; Bouvier-Alias, Magali; Pouget, Noelle; Brevot-Lutton, Ophelie; Bourliere, Marc; Zoulim, Fabien; Plumas, Joel; Aspord, Caroline; Chemin, Isabelle A (27 June 2016). "Pegylated Interferon α-2a Triggers NK-Cell Functionality and Specific T-Cell Responses in Patients with Chronic HBV Infection without HBsAg Seroconversion". PLOS ONE. 11 (6): e0158297. Bibcode:2016PLoSO..1158297B. doi:10.1371/journal.pone.0158297. PMC 4922676. PMID 27348813.
- ^ a b c Poordad, Fred; Landis, Charles S.; Asatryan, Armen; Jackson, Daniel F.; Ng, Teresa I.; Fu, Bo; Lin, Chih-Wei; Yao, Betty; Kort, Jens (August 2016). "High antiviral activity of NS5A inhibitor ABT-530 with paritaprevir/ritonavir and ribavirin against hepatitis C virus genotype 3 infection". Liver International. 36 (8): 1125–1132. doi:10.1111/liv.13067. PMC 5067610. PMID 26778412.
- ^ Gaudieri, Silvana; Rauch, Andri; Pfafferott, Katja; Barnes, Eleanor; Cheng, Wendy; McCaughan, Geoff; Shackel, Nick; Jeffrey, Gary P.; Mollison, Lindsay; Baker, Ross; Furrer, Hansjakob; Günthard, Huldrych F.; Freitas, Elizabeth; Humphreys, Isla; Klenerman, Paul; Mallal, Simon; James, Ian; Roberts, Stuart; Nolan, David; Lucas, Michaela (April 2009). "Hepatitis C virus drug resistance and immune-driven adaptations: Relevance to new antiviral therapy". Hepatology. 49 (4): 1069–1082. doi:10.1002/hep.22773. PMID 19263475.
- ^ Archer, Melissa; Steinvoort, Carin; Oderda, Gary. "UPDATE: New Hepatitis C Combination Agents" (PDF). Utah Department of Health. Archived from the original (PDF) on 27 December 2016. Retrieved 8 September 2016.
- ^ Blight, K. J. (8 December 2000). "Efficient Initiation of HCV RNA Replication in Cell Culture". Science. 290 (5498): 1972–1974. Bibcode:2000Sci...290.1972B. doi:10.1126/science.290.5498.1972. PMID 11110665.
- ^ a b Conte, Immacolata; Giuliano, Claudio; Ercolani, Caterina; Narjes, Frank; Koch, Uwe; Rowley, Michael; Altamura, Sergio; Francesco, Raffaele De; Neddermann, Petra; Migliaccio, Giovanni; Stansfield, Ian (March 2009). "Synthesis and SAR of piperazinyl-N-phenylbenzamides as inhibitors of hepatitis C virus RNA replication in cell culture". Bioorganic & Medicinal Chemistry Letters. 19 (6): 1779–1783. doi:10.1016/j.bmcl.2009.01.066. PMID 19216075.
- ^ Lohmann, V. (2 July 1999). "Replication of Subgenomic Hepatitis C Virus RNAs in a Hepatoma Cell Line". Science. 285 (5424): 110–113. doi:10.1126/science.285.5424.110. PMID 10390360.
- ^ Lohmann, V.; Hoffmann, S.; Herian, U.; Penin, F.; Bartenschlager, R. (1 March 2003). "Viral and Cellular Determinants of Hepatitis C Virus RNA Replication in Cell Culture". Journal of Virology. 77 (5): 3007–3019. doi:10.1128/JVI.77.5.3007-3019.2003. PMC 149776. PMID 12584326.
- ^ Poole, Raewyn M. (13 August 2014). "Daclatasvir + Asunaprevir: First Global Approval". Drugs. 74 (13): 1559–1571. doi:10.1007/s40265-014-0279-4. PMID 25117197. S2CID 207488399.
- ^ "FDA approves new treatment for chronic hepatitis C genotype 3 infections". FDA. 27 July 2015. Archived from the original on July 27, 2015. Retrieved 29 September 2016.
- ^ a b c d e f g h Link, John O.; Taylor, James G.; Xu, Lianhong; Mitchell, Michael; Guo, Hongyan; Liu, Hongtao; Kato, Darryl; Kirschberg, Thorsten; Sun, Jianyu; Squires, Neil; Parrish, Jay; Keller, Terry; Yang, Zheng-Yu; Yang, Chris; Matles, Mike; Wang, Yujin; Wang, Kelly; Cheng, Guofeng; Tian, Yang; Mogalian, Erik; Mondou, Elsa; Cornpropst, Melanie; Perry, Jason; Desai, Manoj C. (13 March 2014). "Discovery of Ledipasvir (GS-5885): A Potent, Once-Daily Oral NS5A Inhibitor for the Treatment of Hepatitis C Virus Infection". Journal of Medicinal Chemistry. 57 (5): 2033–2046. doi:10.1021/jm401499g. PMID 24320933.
- ^ Keating, Gillian M. (3 April 2015). "Ledipasvir/Sofosbuvir: A Review of Its Use in Chronic Hepatitis C". Drugs. 75 (6): 675–685. doi:10.1007/s40265-015-0381-2. PMID 25837989. S2CID 31943736.
- ^ Nakamoto, Shingo (2014). "Hepatitis C virus NS5A inhibitors and drug resistance mutations". World Journal of Gastroenterology. 20 (11): 2902–12. doi:10.3748/wjg.v20.i11.2902. PMC 3961994. PMID 24659881.
- ^ Boucle, Sebastien; Tao, Sijia; Amblard, Franck; Stanton, Richard A.; Nettles, James H.; Li, Chengwei; McBrayer, Tamara R.; Whitaker, Tony; Coats, Steven J.; Schinazi, Raymond F. (September 2015). "Design, synthesis and evaluation of novel anti-HCV molecules that deliver intracellularly three highly potent NS5A inhibitors". Bioorganic & Medicinal Chemistry Letters. 25 (17): 3711–3715. doi:10.1016/j.bmcl.2015.06.031. PMC 4538959. PMID 26099532.
- ^ a b Romine, Jeffrey L.; St. Laurent, Denis R.; Leet, John E.; Martin, Scott W.; Serrano-Wu, Michael H.; Yang, Fukang; Gao, Min; O’Boyle, Donald R; Lemm, Julie A.; Sun, Jin-Hua; Nower, Peter T.; Huang, Xiaohua (Stella); Deshpande, Milind S.; Meanwell, Nicholas A.; Snyder, Lawrence B. (10 March 2011). "Inhibitors of HCV NS5A: From Iminothiazolidinones to Symmetrical Stilbenes". ACS Medicinal Chemistry Letters. 2 (3): 224–229. doi:10.1021/ml1002647. PMC 4017990. PMID 24900306.
- ^ Shi, Junxing; Zhou, Longhu; Amblard, Franck; Bobeck, Drew R.; Zhang, Hongwang; Liu, Peng; Bondada, Lavanya; McBrayer, Tamara R.; Tharnish, Phillip M.; Whitaker, Tony; Coats, Steven J.; Schinazi, Raymond F. (May 2012). "Synthesis and biological evaluation of new potent and selective HCV NS5A inhibitors". Bioorganic & Medicinal Chemistry Letters. 22 (10): 3488–3491. doi:10.1016/j.bmcl.2012.03.089. PMC 7732024. PMID 22507961.
- ^ Strahotin, Cristina Simona; Babich, Michael (2012). "Hepatitis C Variability, Patterns of Resistance, and Impact on Therapy". Advances in Virology. 2012: 267483. doi:10.1155/2012/267483. PMC 3407602. PMID 22851970.
- ^ Owens, Christopher M.; Brasher, Bradley B.; Polemeropoulos, Alex; Rhodin, Michael H. J.; McAllister, Nicole; Wong, Kelly A.; Jones, Christopher T.; Jiang, Lijuan; Lin, Kai; Or, Yat Sun (October 2016). "Preclinical and Clinical Resistance Profile of EDP-239, a Novel Hepatitis C Virus NS5A Inhibitor". Antimicrobial Agents and Chemotherapy. 60 (10): 6216–6226. doi:10.1128/AAC.00815-16. PMC 5038316. PMID 27503644.
- ^ a b c Feld, Jordan J.; Foster, Graham R. (October 2016). "Second generation direct-acting antivirals – Do we expect major improvements?". Journal of Hepatology. 65 (1): S130 – S142. doi:10.1016/j.jhep.2016.07.007. PMID 27641983.